polygenic risk score (PRS)
A polygenic risk score (PRS) is an expression of someone's likelihood of having or developing a particular medical condition. It is a genomic prediction method that can be calculated by evaluating information about multiple genetic markers and variants.
The PRS practice is still evolving and is not commonly used by healthcare professionals. There are no formal guidelines for the process and researchers are still working on improving how the scores are generated.
However, the use of polygenic risk scores is increasing in popularity in the biomedical and social science disciplines. Private healthcare and direct-to-consumer companies -- like genetic testing firm 23andMe -- have also commercialized the practice by calculating and selling personal PRSes to customers.
Polygenic risk scores could potentially become an important tool for healthcare professionals. There is hope that PRSes can be used to improve health outcomes by accelerating diagnoses, assisting providers in healthcare decisions and connecting patients to customized treatments.
However, many health experts have reservations about the usefulness of polygenic risk scores. In the case of Type 2 diabetes, for example, excessive body weight may be more highly predictive than a score calculated from genetic data, and health habits may have more influence on development of the disease than heritable risk does.
How is a polygenic risk score used?
The practice of finding polygenic risk scores is currently evolving and the ways of using the scores to create actionable and cost-effective solutions are still being determined.
The PRS practice can only be effectively applied to populations that have been involved in genomic studies. Unfortunately, the majority of research to date has only evaluated individuals of European ancestry. As a result, there is inadequate data about the genomic variants of other populations and their PRSes cannot be easily calculated.
Furthermore, some research has presented ethical uncertainty regarding how polygenic risk scores might be used -- for example, estimating academic success. There is also concern around how the complex and sometimes ambiguous information produced by PRS studies is interpreted.
However, determining a person's predisposition for disease offers major benefits. For example, women in the United States currently begin the breast cancer screening regimen at the age of 50. If younger women can discover their high risk of breast cancer early on, then they can begin the screening regimen earlier and possibly improve their future outcomes.
If patients discover they're at risk of an incurable disease -- such as Alzheimer's disease -- then their PRSes can be used to match them to clinical trials and drugs that may be beneficial.
Polygenic risk scores also have a variety of potential uses when paired with other complementary, existing measures of risk. For example, a 2013 study performed by Samuli Ripatti at the University of Helsinki discovered that combining PRSes with other conventional risk factors for coronary artery disease -- such as high blood pressure and/or body mass index -- advanced the predictions of patients who would develop the disease. The study revealed that the combination of practices could identify a group of high-risk individuals who would have otherwise been considered at intermediate risk.
How to calculate a polygenic risk score
There are a variety of methods for calculating PRS; however, a polygenic risk score is typically calculated by analyzing multiple SNPs at the same time. SNP -- pronounced snip -- stands for single-nucleotide polymorphism. Nucleotides are the building blocks of DNA; there are four types: adenine (A), thymine (T), cytosine (C) or guanine (G).
A SNP is a DNA sequence variation that occurs when a single nucleotide in the genome differs from other paired chromosomes in the individual or from other members of the same species. The genome is the entire genetic material of an organism. Researchers can take this information, put it into a computer and use statistics to predict how an individual's total SNPs contribute to their disease risk.
Researchers have been able to identify genomic variants associated with diseases by comparing the genetic makeup of individuals with and without specific diseases. There can be hundreds, and even thousands, of variants for each disease, but the large amount of available genomic data has enabled researchers to calculate which variants are most common in groups of people with a disease and increase risk prediction.
How to interpret a polygenic risk score
Polygenic risk scores reveal a person's relative risk for a disease. It is relative because the data used to generate the PRS comes from large-scale genomic studies that compare groups of people with the disease to groups without. In other words, a polygenic risk score will tell an individual how their risk compares to that of a person with a different genetic makeup.
A PRS will not reveal a baseline or timeline for the progression of a disease. Furthermore, a high polygenic risk score does not guarantee that an individual will develop the disease; it simply identifies that the individual has a higher genetic liability -- or likelihood -- of becoming ill and should investigate preventative measures.
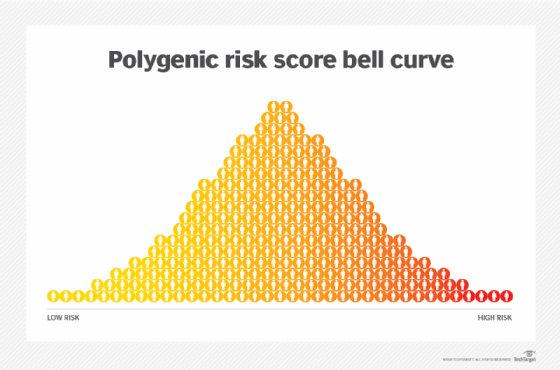
Polygenic risk scores can be displayed on a bell curve. The majority of people will have scores in the middle of the curve; this represents an average risk level for developing a specific disease. Other people will have scores on the tail ends. Scores on the left end indicate a low risk, while scores on the right end indicate a high risk and that individuals should respond with preventative solutions.
Polygenic risk score and 23andMe
At the 2019 SXSW conference, 23andMe announced that it would provide genetics-based risk analysis for Type 2 diabetes to millions of customers. The company correlates genetic information from individuals with health-related data; the resultant data supports the construction of statistical models that can produce a polygenic risk score and predict the likelihood of various traits and conditions from an individual's DNA.
According to the company's news release, 20% of customers would be told that their risk is higher than average; those in the high-risk category would have their scores quantified, as in a one-in-three chance of developing the disease. Those customers would also be targeted with marketing for a health-monitoring app from Lark, a 23andMe partner.